Abstract
The objective of this research was to study the genotype and biotype of BVDV variability among PI dairy cattle in Java, Indonesia between 2016 and 2017. Two hundred isolated buffy coat from dairy cattles that had low reproductive performance and never been vaccinated in Java were used in this studies. Using antigen capture Elisa, 12 out of 200 dairy cattles were positive for the presence of protein Erns BVDV (6.0%). The PI status was confirmed by multiple sequential viral molecular detection. Through phylogenetic and nucleotide sequence analysis of the 5’-Untranslated Region (5’UTR) of the samples investigated, it was determined that all the 12 field positive samples had the BVDV-1 genotype. Three IP-BVDV positive samples (2282-15, 0610-14 and 0813-2) sharing highest similarity (99% homology) with subgenotype BVDV-1c AY763030-1 and KF896608 isolates from Australia. Using Immuno Peroxidase Monolayer Assay (IPMA) the biotype of all the samples were identified as noncytopathic-BVDV. All of the informations would be necessary for designing the diagnostic tool and/or vaccine that match the circulating BVDV subgenotype.
Keywords
Bovine Viral Diarrhea Virus, Genotype, Biotype, 5’-Untranslated Region, Immuno Peroxidase Monolayer Assay
Introduction
Bovine Viral Diarrhea Virus (BVDV), a pestivirus belong to Flaviridae family, is an infectious pathogen affecting cattle in most part of the world [1]. In infected animals, the clinical manifestations caused by BVDV are mainly related to reproductive inefficiency such as poor conception rate, lengthened of calving intervals, congenital malformations, abortions, birth of weak calves and reduction in milk yield resulting significant economic losses to cattle industry worldwide [2, 3]. However, the most devastating consequences is the birth of Persistently Infected (PI) calf which occur when the cows infected by noncytopathic BVDV virus between 30 and 125 days of gestation. Persistently infected calves are immunotolerant and the main source of BVDV transmission in the herd since they shed large amount of the virus throughout their entire lives. Identification of persistent infected calves among cattle population is one of the most important but challenging strategy to control the disease from spreading [4]. Since the persistently infected animals are often born normal and can reach adulthood without any specific clinical signs [1, 5].
Genetically, BVDV is a positive sense single stranded RNA virus of approximately 12.3 kb in length with one Open Reading Frame (ORF) and is flanked on both sides by 5’ and 3’ untranslated regions [6]. Serologically and molecularly, BVDV has been differentiated into two genotypes BVDV-1 and BVDV-2 [7, 8] . Each genotype can be divided into two biotypes, cytopathic (CP) and non-cytopathic based on their cytopathogenecity on cell culture [1, 9] . To date there are many reports on genetic variations of BVDV from many countries [8, 10, 11, 12] but BVDV-1 remains the dominant genotype that has spreading worldwide. The genetic diversity that occur among BVDV isolates is mainly related to the nature characteristic for RNA viruses. According to [13] neither biotype nor genotype are not clinical sign specific. Eventhough the presence of genetic variations of BVDV-1 have not been accepted by the International Committee on Taxonomy of Viruses yet, but they are widely used in molecular epidemiology since the knowledge about the diversity has practical implications to control the occurance of BVDV new variants and to design effective vaccine against the BVDV present in a country [14, 15].
The objective of this study was to study the variation of the genotype and biotype of the BVDV isolated from persistently infected dairy cattle in Java, Indonesia, during 2016–2017, based upon the 5’ untranslated regions. That information is needed for designing the diagnostic tool and/or vaccine that match to the circulating BVDV in Indonesia.
Methods and Materials
Samples
A total of 200 dairy cattle that have low reproductive performance and never been vaccinated were used in this study. The whole blood was withdrawn through coccygeal vein using EDTA-vacutainer tube (Beckton Dickensen) and buffy coat were isolated according to [16]. The presence of the BVDV in the herd was analyzed using antigen capture ELISA (ACE) technique.
Antigen Capture Elisa (ACE)
The presence of protein Erns in the buffy coat were tested individually using Antigen Capture ELISA (IDEXX herdcheck BVDV Antigen Test Kit) as described by the manufacture. Fifty microliters (50 µl) of detection antibodies were dropped into each microwell. Fifty microliters of positive control, negative control and buffy coat samples were added into appropriate well, mixed the content of each well by gently tapping the microplate and incubated for 120 minutes at room temperature. After incubation, empty the liquid in each well followed by washing using 300 µl washed buffer/well for three to five times. One hundred microliters of anti-bovine HRP conjugated were then added into each well and incubated for another 30 minutes at room temperature. After incubation, excess conjugate were removed from microwell by washing the plate three to five times using washed buffer as previously described. One hundred microliters of substrat solution containing 3, 3’, 5, 5’ Tetramethylbenzidine (TMB) were added into each well and the plate was incubated for 10 minutes in the dark at room temperature for color development. The reaction was terminated by adding 100 µl of stop solution into each well. The absorbance of the controls and the samples were measured and recorded at 450 nm in absorbance microplate reader (Bio-Rad Model 680. 2000 Alfred Nobel Drive, Hercules, CA 94547) [17]. The blood samples from the cattles that positive based on ACE test were re-taken 4 weeks later to confirm the persistency of BVDV infection.
RNA Extraction and RT-PCR
The Viral RNA from 200 µl buffy coat suspensions and positive control isolate were extracted using Viral Nucleic Acid Extraction Kit II (Geneaid) as described by the manufacturer instruction. The extracted RNA was subjected to reverse transcription and PCR amplification in one-step reactions using MyTaqTM One-Step RT-PCR Kit (Bioline) according to manufacturer’s specification, in Personal Combi Thermocycler Biometra (37079 Goettingen Germany). Pairs of specific primers for 5’UTR regions of the BVDV genome [18]
(Table 1) were used for amplification of BVDV in the buffy coat. Thermal conditions were as follows: 62°C for 30 min of reverse transcription, 94°C for 2 min of initial denaturing followed by 40 cycles of 94°C for 1 min, 62°C for 1 min, 65°C for 1 min and then final elongation at 65°C for 10 min.
Table 1. Oligonucleotide primers used in this study
|
Primers |
Genotype |
Sequence (5’-3’) |
Product |
References |
|
Forward |
BVDV |
5’ TAG CCA TGC CCT TAG TAG GAC 3’ |
288 bp |
[18] |
|
Reverse |
5’ ACT CCA TGT GCC ATG TAC AGC 3’ |
The BVDV genotyping was done using a nested PCR. In the first cycle of RT-PCR amplification, MyTaqTM One-Step RT-PCR Kit (Bioline) and primer set A (Table 2) was used. The PCR mixtures were then amplified under the following cycling condition: 42°C for 1 h of reverse transcription, 94°C for 3 min of initial denaturing followed by 30 cycles of 94°C for 30 s, 50°C for 45 s, 72°C for 1 min and then one cycle of final elongation at 72°C for 10 min. In the PCR second amplification, a 198 bp DNA product from the first amplication was used as a template for the second round RT-nested PCR. The PCR reaction was prepared similarly to the first amplification but without RT enzyme and primer set B (Table 2). Thermal conditions for the second amplification were as follows: 94°C for 2 min of initial denaturing followed by 30 cycles of 94°C for 30 s, 50°C for 45 s and 72°C for 1 min followed by final elongation at 72°C for 7 min.
Table 2. Oligonucleotide primers for genotyping used in this study
|
Primers |
Genotype |
Sequence (5’-3’) |
Product |
References |
|
Sense Set A |
BVDV-1 and/or BVDV-2 |
5’ GTA GTC GTC AGT GGT TCG 3’ |
198 bp |
[19] |
|
Antisense Set A |
5’ GCC ATG TAC AGC AGA GAT 3’ |
|||
|
Sense Set B |
5’ CGA CAC TCC ATT AGT TGA GG 3’ |
105 bp |
||
|
Antisense Set B |
5’ GTC CAT AAC GCC ACG AAT AG 3’ |
All the PCR products were separated on a 1.5% agarose gel, stained after electrophoresis with ethidium bromide and visualised using ultraviolet transillumination. Two isolates contain Singer strain and 890 strain were respectively used as BVDV type 1 and BVDV type 2 control.
For sequencing, PCR products were purified using High Pure PCR Product Purification Kit (Roche Life Science, Mannheim, Germany). Forward and reverse 5’-UTR sequences for each sampel were aligned and used in phylogenetic analysis. The sequences were compared to other previously publlished sequences. The sequence identities of nucleotide, as well as the estimation of the evolutionary divergence between sequences were analysed using DNA-Baser and Mega7 software, respectively [20]. The same tool was used to perform Neighbor-Joining analysis.
Immunoperoxidase Monolayer Assay (IPMA)
Immuno peroxidase monolayer assay were used for determination of BVDV biotype. Microtitration flat bottom plate wells were seeded with 100, 000 cells/ml Madin-Darby bovine kidney (MDBK) cells and incubated at 37°C in 5% CO2 atmosphere. After 24 hours, MDBK cells in every well were inoculated with 50 µl of the positive isolates. For non-infected control, the MDBK cells in the microtitration plate well was added with 50 µl distilled water. The MDBK infected and non-infected cells were incubated for another 72-hours at 37°C in 5% CO2 atmosphere. After incubation, the microtitration plates were drained, rinsed three times with wash buffer, and fixed with fixing buffer containing 35% acetone in phosphate-buffered saline (PBS, pH 7, 4) and 0, 02% bovine serum albumin for 10 minutes at room temperature. After fixation the cells were then incubated with 50 µl blocking serum solution for 10 minutes. After the incubation, the remnant of blocking serum solution left on the well was drained off. Fifty microliters monoclonal antibody anti BVDV 15c5 (1: 100 dilution) were added onto infected and non-infected MDBK cells and incubated for 60 minutes at room temperature. After incubation, the microtitration plate was drained and then rinsed 3 times with wash buffer (PBS solution containing 0.05% Tween 20) of 2 minutes each and drained off. Into each well was then added 50 µl with a secondary antibody solution-labeled with biotin and incubated for 10 minutes at room temperature. After incubation, the microtitration plate was drained and then rinsed 3 times with wash buffer 2 minutes each and drained off. Fifty microliters of streptavidin peroxidase conjugated solution were added into each microtitration well followed by another incubation step for 15 minutes. Following the incubation, the microtitration plate was drained and then rinsed 2 times 1 minute each with wash buffer. Fifty microliters of the mixed-substrate (H2O2) chromogen (3.3’-diaminobenzidine) solution were added into each well and left to react with the cells for 1 hour. The enzymatic reaction was stopped by rinsing the microtitration plate with tap water, drained off and followed by counterstaining the cells with hematoxylin for 3 minutes. The cells were then examined under light microscope [21].
Results
In this study, using antigen capture Elisa, 12 out of 200 dairy cattles were positive for the presence of protein Erns BVDV (6.0%). For confirming the virus persistence, blood samples from BVDV positive animals were re-taken and re-tested one month apart using RT-PCR technique. The results show specific product at 288 bp visible on the 1.5% agarose gel (Figure 1).
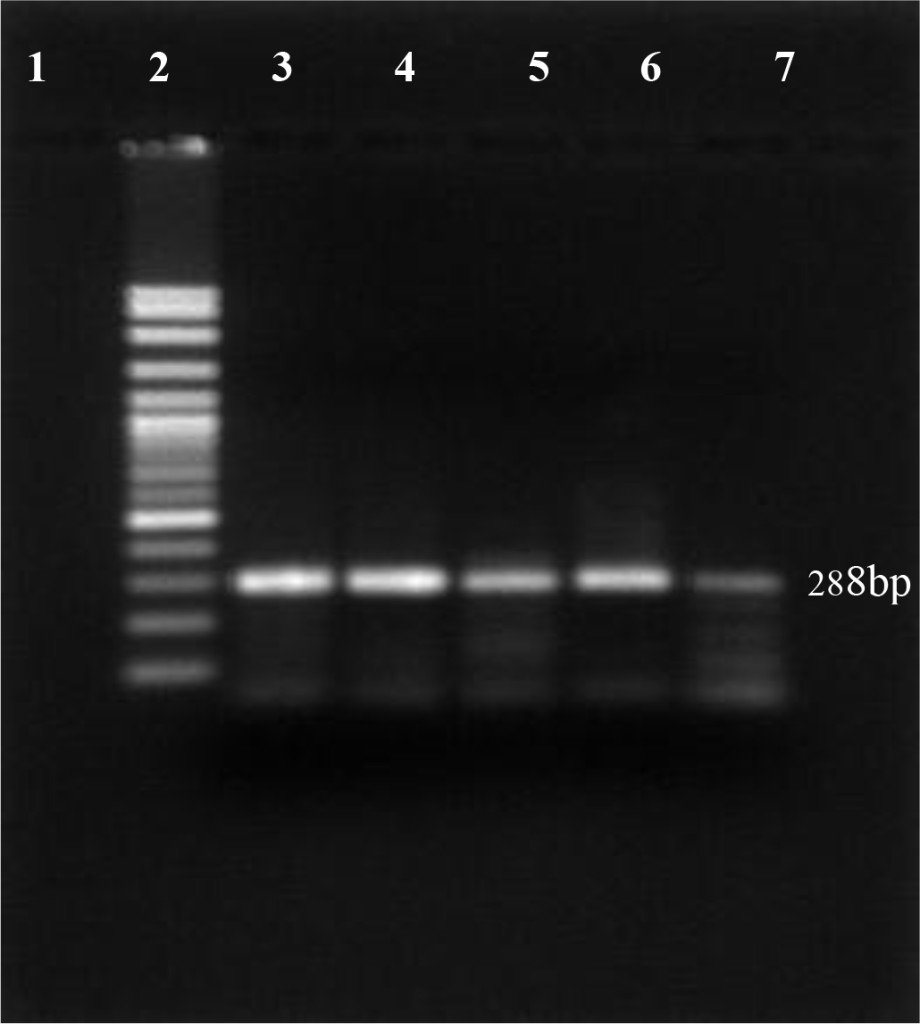
Figure 1. PI-BVDV detection by RT-PCR conventional technique. (Lane 1: negative control, lane 2: DNA marker 100 bp, lane 3: positive control, lane 4 – 7 PI-BVDV positive from field samples).
Using the oligonucleotide primers pairs listed in Table 2, after the first amplification, the DNA bands from all field samples and positive control either BVDV-1 and BVDV-2 were clearly visible at 198 bp (Figure 2). After the second amplification, the BVDV genotype 1 were no longer visible but BVDV genotype 2 should be clearly seen at 105 bp. In this study, however, only 890 strain (BVDV-2 positive control) showed positive result (Figure 3). Based on the sequence analysis, all of the positive samples in this study were clustered within the BVDV-1 genotype and no evidence for the presence of BVDV genotype 2.
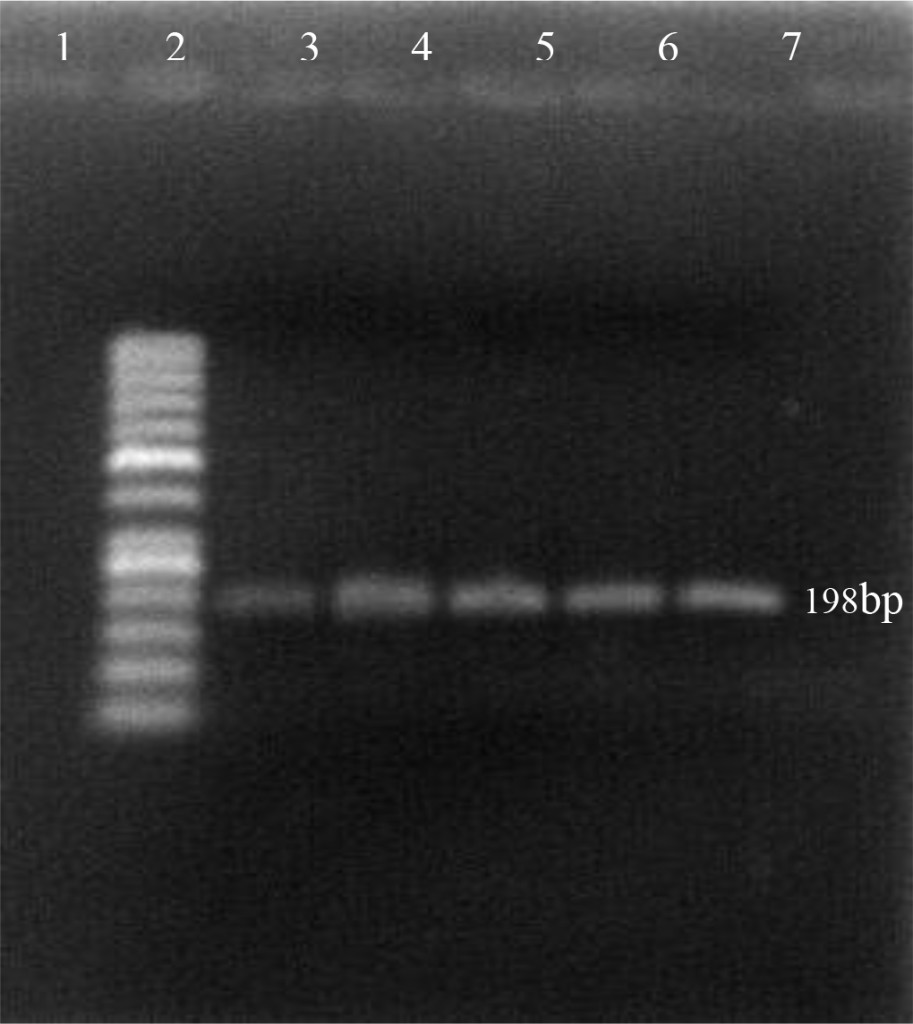
Figure 2. First PCR amplification of RNA extracted from positive PI-BVDV samples. (Lane 1: negative control, lane 2: DNA marker 50 bp, lane 3: Singer strain BVDV-1 positive control, lane 4–7: BVDV-1 positive from field samples).
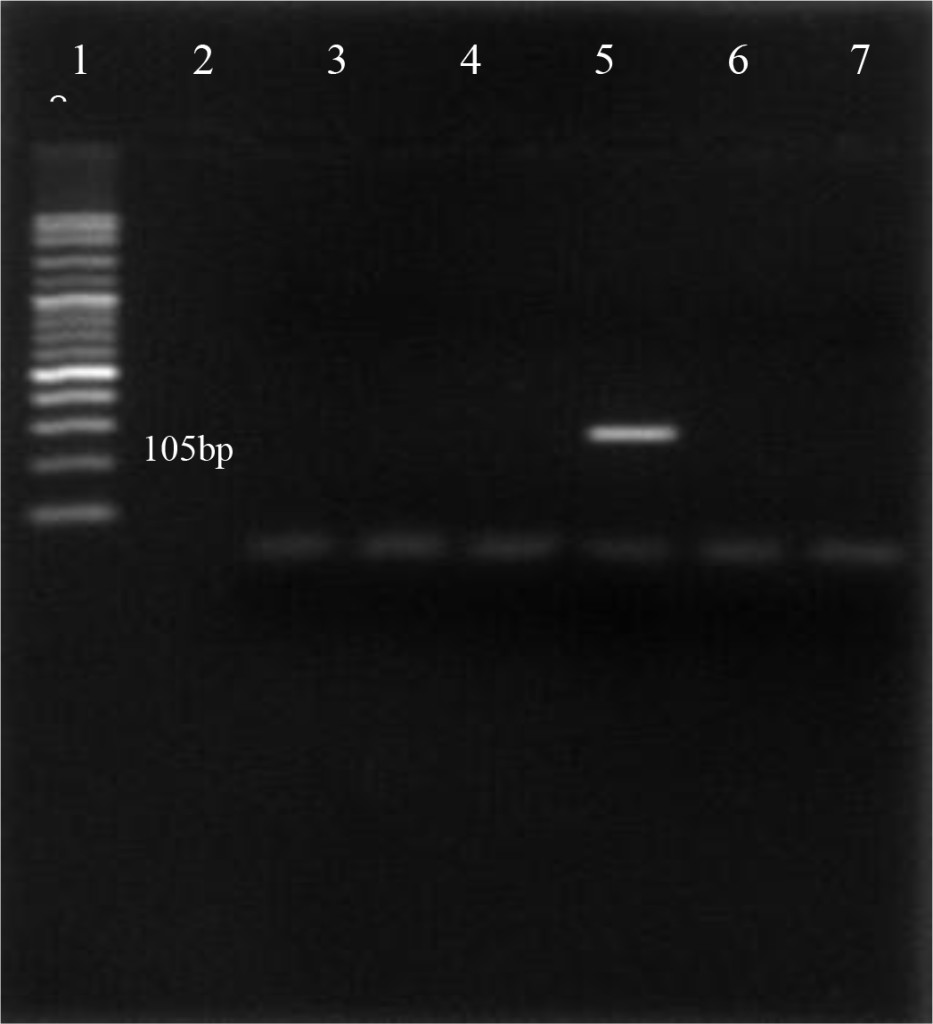
Figure 3. First PCR amplification of RNA extracted from positive PI-BVDV samples. (Lane 1: DNA marker 50 bp, lane 2: negative control, lane 3: Singer strain BVDV-1 positive control, lane 4, 5, 7, 8: BVDV-1 positive field samples, lane 6: 890 strain BVDV-2 positive control).
Discussion
Of the 200 animals tested 12 (6%) were positive for persistently infected BVDV (PI- BVDV). Regardless of their prevalence, PI-BVDV animals are the main source of infection within the herd [22]. The generation of persistently infected animals is more often when BVDV infection occur in-utero between 45 to 130 days of gestation period. Many PI-BVDV animals can be clinically healthy, although their life expectancy is low (< 2 years). However, according to [23]. PI-BVDV animals that can live beyond the age of 2 years were possibly related to the nonpathogenecity of the virus. The age range of the positive PI-BVDV animals in this study is between 2 weeks – 36 months. According to [16] and [24] in close confinement housing operation a PI animal can infect up 90% of the herd regardless of their prevalence. Results of this study have verified how well the persitently infected animals in a dairy herd with no BVDV vaccination had infected cattle in the same herd and confirm the magnitude of the infection in concentrate cattle production system.
The genetic variations of the PI-BVDV field positive samples in this study was done by determining the nucleotide sequences of the 5’UTR region that are highly conserved among all of the pestiviruses. High conservation permits rapid and accurate acquisition of the sequence data but may not be good target for interfering phylogeny [25, 26]. Based on the result, the genetic typing of viral RNA revealed that among PI-BVDV positive animals all were BVDV genotype 1 (BVDV-1). The result is in agreement with previous research finding done by [27] and [28]. BVDV-1 is distributed more widely throughout the world compared to BVDV-2 [26]. BVDV-2 was first reported in the USA but later has been described in European and Asian countries at lower rate than BVDV-1 [29, 30, 31].
Based on the phylogenetic analysis (Figure 4, Table 3), 9 of the samples clustered within the BVDV-1a subgenotype whereas 3 other samples belong to the BVDV-1c subgenotype. The result showed that the majority of BVDV subgenotype identified since 2015 in dairy cattle in Java is BVDV-1a [28]. Further result revealed that isolates number 2282–15, 0610–14 and 0813–2 sharing high similarity (98% homology) with the representative subgenotype BVDV-1c Genebank accession number AY763030–1 and KF896608 from Australia. The result is logic since the majority of dairy cattle used in this study were historically coming from Australia in which the BVDV-1c sub-genotype is predominant [32]. Using different genomic regions (NS5B) and larger samples (588) tested for genotyping of BVDV, [27] had found 4 BVDV-1 subgenotypes (1a, 1b, 1c, 1d) in cattle in Java. BVDV variability were generally analyzed using different genomic regions such as 5’UTR [33], NPRO [34], E2 [35], NS2–3 [36] and NS5B [37] and the result is not comparable to each other. Time periods for sampling also has an effect on the result of BVDV subgenotyping [28].Factor that influence the spread of the BVDV sub genotypes is unclear and do not appear to be affected by vaccine used but rather in part reflect the antigen diversity between strains [38]. According to [29], the genetic variation of the BVDV in a given geographic area has been largely influenced by animal movement within countries and/or introduction from other countries.
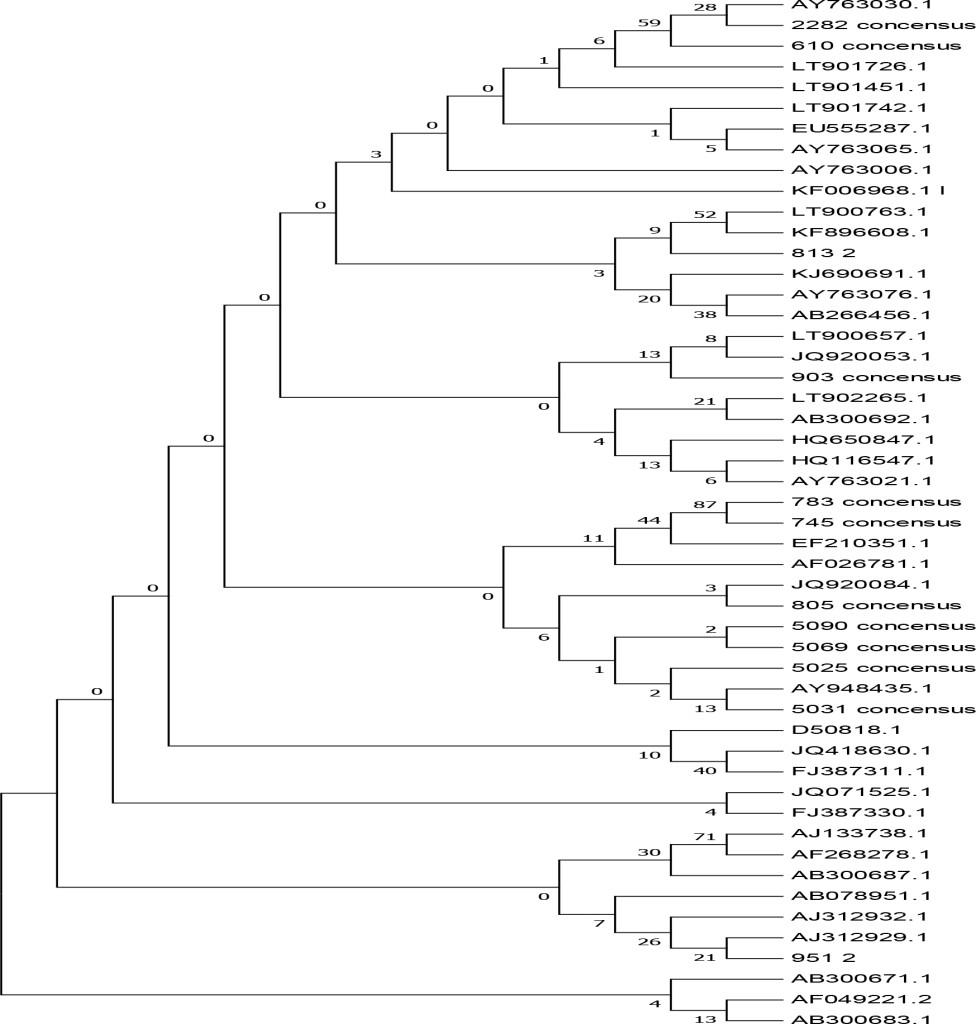

Figure 4. Phylogenetic tree of the 5’ untranslated regions (UTR) of BVDV strains and isolates. The tree was generated from comparative alignment of sequences from 288bp of the 5’UTR of the BVDV genome,
Table 3. Summary of genomic form of BVDV isolated from persistently infected dairy cattle in this study
|
No. |
Isolate # |
BVDV Biotype |
BVDV Genotype |
BVDV subgenotype |
|
1. |
JT-2282–15 |
NCP** |
Type 1 |
1-c |
|
2. |
JT-0610–14 |
NCP |
Type 1 |
1-c |
|
3. |
0813–2 |
NCP |
Type 1 |
1-c |
|
4. |
0903–16 |
NCP |
Type 1 |
1-a |
|
5. |
0783–16 |
NCP |
Type 1 |
1-a |
|
6. |
0745–16 |
NCP |
Type 1 |
1-a |
|
7. |
0805–14 |
NCP |
Type 1 |
1-a |
|
8. |
5090–16 |
NCP |
Type 1 |
1-a |
|
9. |
5069–17 |
NCP |
Type 1 |
1-a |
|
10. |
5025–17 |
NCP |
Type 1 |
1-a |
|
11. |
5031–17 |
NCP |
Type 1 |
1-a |
|
12. |
0951–2 |
NCP |
Type 1 |
1-a |
**NCP=non-cytopathic
In this study, staining using IPMA technique was performed on MDBK cells that had infected with BVDV isolates obtained from cows with persistent BVDV infections. Positive results were characterized by brownish precipitates in single-layer cells (Figure 5). In the cells culture, within 24–48 hours after inoculation the virus did not cause vacuolization in the cytoplasm and / or cell damage. This suggests that the biotype of IP-BVDV in this study was non-cytopathic (NCP-BVDV) and proved that PI animals only have NCP-BVDV biotype. The NCP biotype is mostly occur in virus that transmitted vertically and is considered as a marker for persistentcy of BVDV in the herd [39]. Cytopathology in tissue culture does not correlate with virulence of the virus in vivo. In another word biotype is not clinical manifestation specific [13, 40]. Based on epidemiologic studies, NCP-BVDV is a more common biotype than BVDV cytopathic [39] .
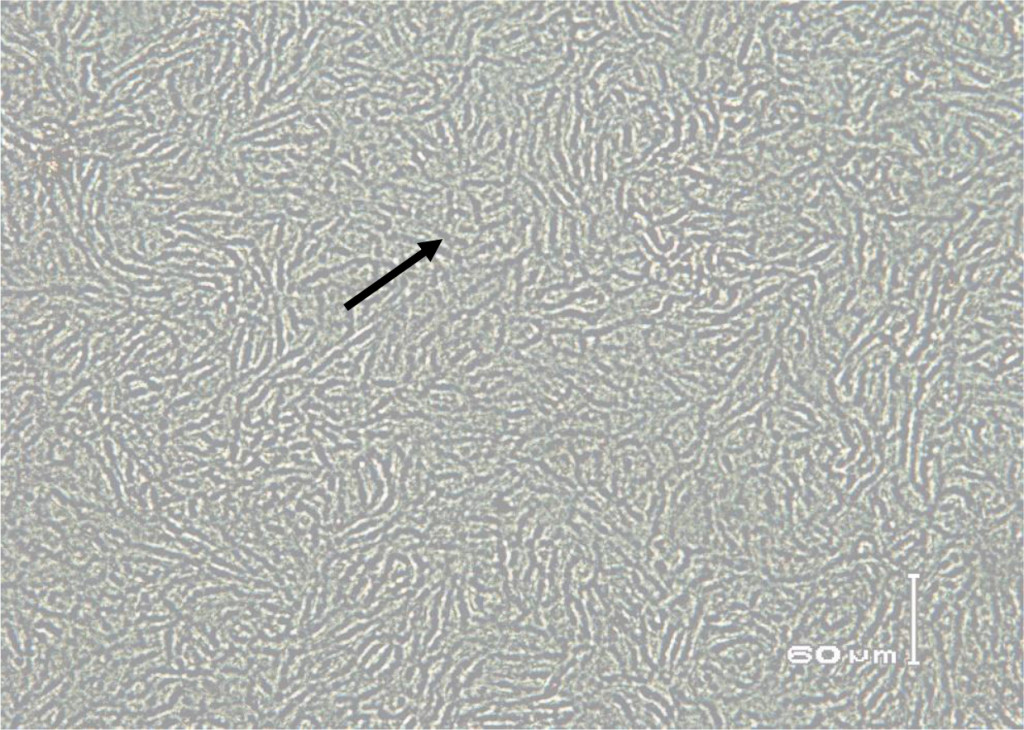
Figure 5. Immunoperoxidase monolayer assay (IPMA) in Madin Darby Bovine Kidney (MDBK) immune cells infected with serum that does not contain BVD virus (negative control).
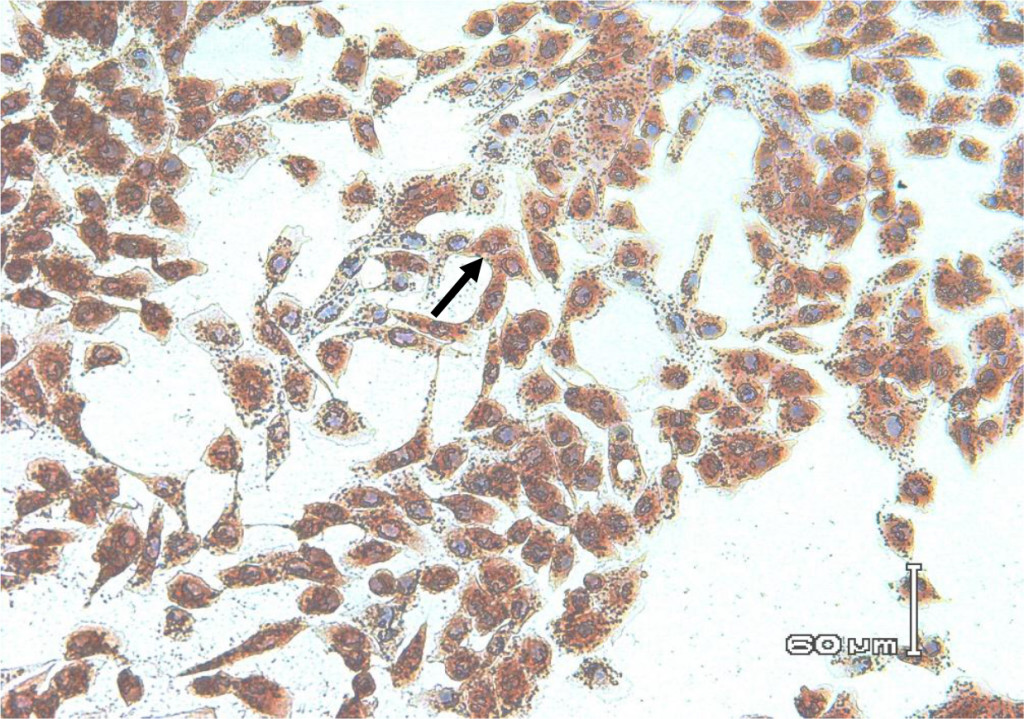
Figure 6. Biocytes of MDBK cells infected with non-cytopathic BVDV field isolates for 72 hours. Using IPMA cell-positive staining infected with BVDV appears brownish in the cytoplasm and nucleus.
The results of the study confirm the presence of the persistently- infected BVDV dairy cattles and provide information about BVDV biotypes and subgenotypes that are dominant among PI-BVDV positive dairy catttles, in Java, Indonesia between 2016 and 2017. All of the informations will be necessary for designing the diagnostic tool and/or a vaccine that match the circulating BVDV subgenotype in Indonesia in the future.
Author’s contribution
Sugiyono played a valuable role in blood sampling and preparing the buffy-coat from the whole blood for further analyses. Raden Wasito had responsibility in doing the IPMA. Hastari Wuryastuti had role in doing the RNA extraction, PCR analysis and sequencing. All researchers participated equally in preparing the manuscripts for journal publication.
Acknowledgment
The authors greatly acknowledge to the Direktorat Riset dan Pengabdian Masyarakat Direktorat Jenderal Penguatan Riset dan Pengembangan, Kementrian Riset, Teknologi, dan Pendidikan Tinggi and Gadjah Mada University for financial support through Penelitian Dasar Unggulan Perguruan Tinggi, Gadjah Mada University No.195/UN1/DITLIT/DIT-LIT/LT/2018 Tanggal 5 Maret, 2018. The authors are deeply grateful for the thoughtful advice given by Prof. Dr. Roger K. Maes, Michigan State University, East Lansing, MI, USA during the preparation of this manuscript.
References
- Brodersen BW (2014) Bovine Viral Diarrhea Virus infections: Manifestations of infection and recent advances in understanding pathogenesis and control. Vet Pathol 51: 453–464.
- Heuer C, Healy A, Zerbini C (2007) Economic effects of exposure to bovine viral diarrhea virus on dairy herds in New Zealand. J Dairy Sci 90: 5428–5438. [crossref]
- Richter V, Lebl K, Baumgartner W, Obritzhauser W, Kasbohrer A, Pinior B, et al (2017) A systematic worldwide review of the direct monetary losses in cattle due to bovine viral diarrhea virus infection. Vet J 220: 80–87.
- Stalder H, Hug C, Zanoni R, Vogt HR, Peterhans E, Schweizer M, Bachofen C, et al (2016) A nationwide database linking information on the hosts with sequence data of their virus strains: A useful tool for the eradication of bovine viral diarrhea (bvd) in Switzerland. Virus Res 218: 49–56.
- Thomann B, Tschopp A, Magouras I, Meylan M, Schupbach-Regula G, Hasler B, et al (2017) Economic evaluation of the eradication program for bovine viral diarrhea in the Swiss dairy sector. Prev Vet Med 145: 1–6.
- Meyers G, Thiel HJ (1996) Molecular characterization of pestiviruses. Adv Virus Res 47: 53–118. [crossref]
- Ridpath JF (2003) BVDV genotypes and biotypes: practical implications for diagnosis and control. Biologicals 31: 127–131. [crossref]
- Yesilbag K, Alpay G, Becher P, et al (2017) Variability and global distribution of subgenotypes of bovine viral diarrhea virus. Viruses 9: 128–146.
- Ammari M, McCarthy FM, Nanduri B, et al (2012) Understanding the pathogenesis of cytopathic and noncytopathic bovine viral diarrhea virus infection using proteomics. In: Heazlewood J (Ed.). Proteomics Application in Biology. In Tech Pg No: 53–66.
- Vilcek S, Paton DJ, Durkovic B, Strojny L, Ibata G, et al (2001) Bovine viral diarrhoea virus genotype 1 can be separated into at leasteleven genetic groups. Arch Virol 146: 99–115.
- Zhang J-J, Ma C, Li X-Z, Liu Y-X, Huang K, et al (2013) Genetic diversity of bovine viral diarrhoea virus in beijing region, China from 2009 to 2010. Afr J Microbiol Res 7: 4934–4939.
- Bazzucchi M, Bertolotti L, Giammarioli M, Casciari C, Rossi E, Rosati S, DeMia MG, et al (2017) Complete genome sequence of a bovine viral diarrhea virus subgenotype 1h strain isolated in Italy. Am Soc Microbiol 5: 1697–1716.
- Ahn BC, Walz PH, Kennedy GA, Kapil S, et al (2005) Biotype, genotype, and clinical presentation associated with bovine viral diarrhea virus (BVDV) isolates from cattle. Intern. J Appl Res Vet Med 3: 319–325.
- Simmonds P, Becher P, Collett MS, Gould EA, Heinz FX, et al (2012) Family Flaviridae. In: King, A.M.Q., Adams MJ, Carston EB, Lefkowitz EJ (Eds.). Virus Taxonomy Ninth Report of the International Committee on Taxonomy of Viruses. Elsevier Academic Press, Amsterdam, Boston, Heidelberg Pg No: 1003–1020.
- Moennig V, and Becher P (2018) Control of bovine viral diarrhea. Pathogens 7: 29–40.
- Peddireddi L, Foster KA, Poulsen EG, An B, Hoang QH, et al (2018) Molecular detection and characterization of transient bovine viral diarrhea virus (BVDV) infections in cattle commingled with ten BVDV persistently infected cattle. J Ve. Diag Invest : 1–10.
- Wuryastuti H, Putro PP, Wasito R, et al (2014) Identifikasi, isolasi dan propagasi virus penyebab infeksi persisten bovine viral diarrhea pada sapi untuk produksi isolat lokal unggulan. Laporan Penelitian Unggulan Perguruan Tinggi.Tahun Pertama. Universitas Gadjah Mada 2014.
- Weinstock D, Bhudevi B, Castro AE, et al (2001) Single-tube single-enzyme reverse transcriptase PCR assay for detection of bovine viral diarrhea virus in pooled bovine serum. J Clin Microbiol 39: 343–346.
- El-Kholy AA, Bolin SR, Ridpath JF, Arab RMH, Abou-Zeid AA, et al (1998) Use of polymerase chain reaction to simultaneously detect and type bovine viral diarrhea viruses isolated from clinical specimens. Rev Sci Tech Off Int Epiz 17: 733–742.
- Kumar S, Stetcher G. Tamura K, et al (2016) Mega7: Molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 3: 1870–4.
- Wuryastuti H, and Wasito R. Application of the immunoperoxidase monolayer assay to characterize non-cytopathic bovine viral diarrhea virus of Indonesian samples. Indon J Biotech. 1997: 154–158.
- Fulton RW (2015) Impact of species and subgenotypes of bovine viral diarrhea virus on control by vaccination. Anim Health Res Rev 16: 40–54. [crossref]
- El-Bogoury GF, El-Nahas EM, Khadr KA, Ali NM, et al (2014) Isolation and genotyping of bovine viral diarrhea virus: Field isolates from infected cattle in Kaluobia during 2011. BVMJ 23: 6–10.
- Houe H (1999) Epidemiological features and economical importance of bovine viral diarrhea virus (BVDV) infections. Vet Microbiol 64: 89–107.
- Bedekovic T, Lojkic I, Lemo N, CAC Z, Cvetnic Z et al (2012) Genetic typing of Croation bovine viral diarrhea virus isolates. Vet Archiv 82: 449–462.
- Soltan MA, Wilkes RP, Elsheery MN, Elhaig MM, Riley MC, et al (2015) Circulation of bovine viral diarrhea virus-1 (BVDV-1) in dairy cattle and buffalo farms in Ismailia Province, Egypt. J Infec Dev Ctries 9: 1331–1337.
- Saepulloh M, and Sendow I (2015) Identification and characterization of bovine viral diarrhea virus from Indonesian cattle. J Vet 16: 1–7.
- Wuryastuti H, Putro PP, Wasito R, Maes, RK, et al (2015) Genetic variability of Bovine Viral Diarrhea Virus in persistently infected cattle in Central Java, Indonesia. Merit Res J Microbiol Biol Sci 3: 031–036.
- Luzzago C, Lauzi S, Ebranati E, Giammarioli M, Moreno A, et al (2014) Extended genetic diversity of bovine viral diarrhea virus and frequency of genotypes and subtypes in cattle in Italy between 1995 and 2013. Bio Med Res Pg No: 147–151.
- Fulton RW, Whitley EM, Johnson BJ, Ridpath JF, Kapil S, et al (2006) Prevalence of bovine viral diarrhea virus (BVDV) in persistently infected cattle and BVDV subtypes in affected catle in beef herds in south central United States. Can. J. Vet Res 73: 283–291.
- Jackova A, Novackova M, Pelletier C, Audeval C, Gueneau E, et al (2008) The extended genetic diversity of BVDV-1: typing of BVDV isolates from France. Vet Res Commun 32: 7–11. [crossref]
- Hay KE, Ambrose RC, Morton JM, Horwood PF, Gravel JL, et al (2016) Effects of exposure to bovine viral diarrhea virus 1 on risk of bovine respiratory disease in Australian feedlot cattle. Prev Vet Med 126: 159–169.
- Abe Y, Tamura T, Torii S, Wakamori S, Nagai M, et al (2016) Genetic and antigenic characterization of bovine viral diarrhea viruses isolated from cattle in Hokkaido, Japan. J Vet Med Sci 781: 61–70.
- Ridpath JF, Fulton RW, Kirkland P, Neil JD, et al (2010) Prevalence and antigenic differences observed between bovine viral diarrhea virus subgenotypes isolated from cattle in Australia and feedlots in southwestern United States. J Vet Diagn Invest 22: 184–191.
- Silveira S, Weber MN, Mosena AC, daSilva MS, Streck AF, et al (2015) Genetic diversity of Brazilian Bovine Pestivirus detected between 1995 and 2014. Transbound Emerg Dis 64: 613–623.
- Xia H, Liu L, Wahlberg N, Baule C, Belak S, et al (2007) Molecular phylogenetic analysis of bovine viral diarrhea virus, a Bayesian approach. Virus Res 130: 53–62.
- Mishra N, Dubey R, Rajukumar K, Tosh C, Tiwari A, et al (2007) Genetic and antigenic characterization of bovine viral diarrhea virus type 2 isolated from Indian goats (Capra hircus). Vet Microbiol 124: 340–347.
- Ambrose RK, Gravel JL, Commins MA, Fowler EV, Mahony TJ (2018) In Vivo Characterisation of Five Strains of Bovine Viral Diarrhoea Virus 1 (Subgenotype 1c). Pathogens 7: E12 [crossref]
- Bazid AI, Nayel MA, Elsify AM, Zaghloul AH, Khalil SA, et al (2015) Molecular diagnosis and genotyping of Bovine viral diarrhea virus. Minufiya Vet J 9: 1–13.
- Ridpath JF (2005) Practical significance of heterogeneity among BVDV strains: Impact of biotype and genotype on U.S. control programs. Prev Vet Med 72: 17–30.