Abstract
Background: The Qianghuo herb pair is renowned in traditional Chinese medicine (TCM) for its blood-activating and stasis-resolving properties, frequently incorporated into formulas for leukemia treatment. However, the specific relationship between the Qianghuo herb pair and leukemia, along with its underlying mechanisms, remains elusive. Based on network pharmacology and molecular docking technology, explore the relevant targets of Qianghuo and its mechanisms in treating leukemia.
Method: Diverse databases were utilized to investigate the chemical components and their respective targets of Qianghuo, alongside acquiring disease-related target information. A network relationship diagram of Qianghuo’s active ingredients, targets, and diseases was constructed using Cytoscape 3.8.2 software. Protein-protein interaction (PPI) networks were created and potential pathways were explored through Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) analysis. Molecular docking was utilized to assess the interactions between active ingredients and potential targets.
Result: In Qianghuo, a total of 15 active ingredients and 554 corresponding targets were obtained. The main active ingredients include Ammidin, 6’-Feruloylnodakenin, 8-geranoxy-5-methoxypsoralen, coumarin, glycoside, Demethylfuropinnarin, Bergaptin, etc. Among them, 162 targets intersect with drug-disease interactions, with AKT1, EGFR, SRC, CASP3, and HSP90AAA1 identified as core targets. Functional enrichment analysis of Gene Ontology (GO) yielded 870 entries, while Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis identified 154 pathways, notably including Pathways in cancer, PI3K-Akt signaling pathway, and Human papillomavirus infection. Molecular docking simulations revealed significant binding affinity between the main active ingredient of the drug and the core target proteins.
Conclusion: This study successfully elucidates the mechanism of action of Qianghuo in treating leukemia, predicts the potential targets of traditional Chinese medicine, elucidates complex signaling pathways, and demonstrates its therapeutic characteristics of multi-components, multi-targets, and multi-pathways. This provides a basis for exploring alternative approaches to researching novel leukemia drugs at the level of network pharmacology.
Keywords
Qianghuo, Leukemia, Network pharmacology, Bioinformatics, Molecular docking
Introduction
Leukemia is a malignant hematological disease with a high incidence in our country, which usually occurs in children and adolescents [1]. It is categorized into four types based on cell differentiation and disease progression: chronic lymphocytic leukemia (CLL), chronic myeloid leukemia (CML), acute lymphoblastic leukemia (ALL), and acute myeloid leukemia (AML). Among them, acute myeloid leukemia stands as the most prevalent, constituting approximately 70% of adult leukemias. This disease develops rapidly, with its natural course typically lasting only a few months. Studies indicate a close association between leukemia and various factors such as genetics, smoking, obesity, alcoholism, exposure to chemical agents (like benzene, formaldehyde, etc.), high-dose radiation, and contact with electromagnetic fields [2-4]. Currently, leukemia treatment methods encompass chemotherapy, immunotherapy, targeted therapy, etc. However, owing to the clonal biological traits inherent in leukemia, the cure rate remains low. Prolonged chemotherapy often induces a sharp decline in the patient’s physical resilience, potentially leading to complications such as bleeding and infection. Hence, the pursuit of novel leukemia drugs with potent efficacy at affordable costs has emerged as a focal point in drug research and development. Traditional Chinese medicine (TCM), with its notable advantages including affordability, minimal side effects, and evident therapeutic effects, has garnered attention as a promising avenue in current drug research and development [5,6].
Qianghuo documented in “Shen Nong’s Materia Medica”, consists of the dried rhizome and root of Qianghuo or Qianghuo Latifolia, belonging to the Umbelliferae family. It holds significance as one of the vital and precious medicinal resources in our country. Qianghuo is mainly composed of sugars, volatile oils, coumarin and other ingredients [7]. The synergy of these components addresses various health issues. It is reported that Qianghuo has anti-inflammatory, analgesic, antipyretic, antibacterial, anti-cancer cell proliferation, anti-arrhythmic, anti-thrombosis and promotion of brain blood circulation effects [8]. Its clinical versatility spans multiple disciplines, with applications in conditions like cancer, cardiovascular and cerebrovascular diseases, rheumatology, and immunology, among others. Despite its widespread clinical use, there remains a gap in understanding the mechanism of Qianghuo in treating leukemia.
Therefore, in order to clarify the effect and mechanism of Qianghuo in the treatment of leukemia, and to expand its clinical application, the potential targets and core active ingredients of Qianghuo anti-leukemia were screened by compound network pharmacology approach in this study. Then, the core active components and the core targets that exert anti-leukemia effects were reverse verified by molecular docking technology. Subsequently, the signaling pathway of Qianghuo in the treatment of leukemia was elucidated, and the therapeutic effect of Qianghuo in leukemia was predicted. These results provide ideas and directions for elucidating the mechanism of Qianghuo in the treatment of leukemia at the molecular level.
Materials and Methods
Software and Databases
Databases used: Drug ingredients and target acquisition database TCMSP (https://tcmspw.com/tcmsp.php), disease target acquisition database Gene Cards (https://www.genecards.org/), OMIM database (https://www.omim.org/), Therapeutic Target Database (http://db.idrblab.net/ttd); Protein information acquisition database (https://www.uniprot.org/); Gene interaction database String (https://www.string-db.org/); Intersection gene acquisition Venny 2.1 (http://bioinfogp.cnb.csic.es/tools/venny/index.html); Software used: Cytoscape V3.8.0 software [9], AutoDock 1.5.7 [10], Openbabel, Pymol [11].
Acquisition of Active Ingredients of Qianghuo and Screening of Targets
With the help of the TCMSP database, the corresponding active ingredients were searched using the keyword “Qianghuo”, and the two indicators of oral bioavailability (OB) ≥ 30% and drug likeness (DL) ≥ 0.18 were set as filtering conditions. Under the Related Targets column, search for target proteins related to the effective active ingredients respectively; then use the Uniprot database and Perl language to standardize the human gene names corresponding to the target proteins.
Obtaining Leukemia Target Proteins
Use OMIM, Genecards, TTD, DisGNET database and other databases, search with “leukemia” as the keyword, select the species as Homo sapiens, and obtain leukemia-related targets. The targets of the database were merged and duplicate targets were removed and corrected by Uniport as the final result of leukemia-related targets for subsequent analysis. Use the WeChat website to draw a Venn diagram of compounds and disease targets.
Construct a Nodule-active Ingredient-Target-Disease Interaction Network and Active Ingredients-Target-Pathway Network Diagram
First, we’ll match the target genes of the active ingredients of Qianghuo obtained from the TCMSP database with leukemia-related target genes. Then, using the WeChat website, we’ll draw a compound-disease target network. The intersection targets will be visualized through a Venn diagram to identify key targets specific to Qianghuo in treating leukemia. Next, we’ll submit the obtained intersection targets to the String11.0 database, setting the biological species as “Homo sapiens” to construct a protein interaction network. Finally, we’ll import the data into Cytoscape 3.7.1 for Network analysis and then screen the core targets based on degree values.
GO Function and KEGG Function Enrichment Analysis
DAVID 6.8 database was used for GO analysis, and the key targets were analyzed in the four modules of Biological process (BP), Cellular component (CC) and Molecular function (MF), and Kyot Encyclopedia of genes and genomes (KEGG). Analysis to identify key targets and enriched pathways.
Molecular Docking of Important Components and Core Targets of Qianghuo
Use the PubChem database to obtain the small molecule structure, save it in sdf format, convert the format in Openbable software, and save it in mol2 format. Download the protein structure of the relevant target through the PDB database, use Pymol software to remove water and ligands, and save it in pdb format. Finally, the database Auto Dock Vina is used to preprocess the small molecule ligands and protein receptors, and its plug-in is used to obtain docking active sites and perform molecular docking to obtain the lowest binding energy for docking between the target protein and the compound. Pymol software is used for visualize the docking results [12].
Results and Discussion
Traditional Chinese herbal medicine serves as a vital reservoir for the development of effective anticancer drugs, offering numerous active molecules with unique structures that can inspire the creation of novel therapeutics. Moreover, the rapid advancements in network pharmacology provide robust support for understanding the mechanisms underlying Chinese herbal medicine’s efficacy in cancer treatment [13]. Qianghuo, as a traditional Chinese medicinal material, has emerged as a research hotspot in recent years, and its compound preparations, action targets, pharmacological activities and other aspects have been continuously explored. Research has demonstrated that Qianghuo contains compounds with varying degrees of inhibitory effects on the proliferation of various cancer cells. This highlights its potential as a promising candidate for further exploration in anticancer drug development. Yang and co-works investigated the two major active ingredients in Qianghuo, nodol and isoimperatorin, and confirmed their inhibitory effects on the proliferation of rat fibroblast-like synovial cells [14]. Notably, the inhibitory effect of isoimperatorin was found to be superior to that of chanzolinol. Chang and co-works discovered the association of the SET domain-containing 2 (SETD2) gene with acute lymphoblastic leukemia [15]. Using this discovery as a starting point, they conducted sub-docking and molecular dynamics simulations. Their findings indicated that coniferin and coniferyl ferulate exhibit high affinity with the SETD 2 protein, demonstrating strong binding affinity and stable interaction. Importantly, coniferyl ferulate can be isolated from Rhizoctonia latifolia. Therefore, the traditional Chinese medicine compound coniferyl februate emerges as a promising lead compound with potential to develop into an anti-leukemia drug. While the aforementioned research findings offer valuable insights into the treatment of leukemia with Qianghuo, much remains to be explored regarding its active ingredients and pharmacological mechanisms. Therefore, further research in this area is necessary to elucidate these aspects and unlock the full therapeutic potential of Qianghuo in leukemia treatment.
Screening of Active Compounds and Targets of Qianghuo
Utilizing the TCMSP database, a total of 185 compounds contained in Qianghuo were collected. According to the ADME standard, which sets two parameter standards of OB≥30% and DL≥0.18, a total of 16 compounds were screened, primarily including Ammidin, 6′-Feruloylnodakenin, Bergaptin, sitosterol, etc., (Table 1 and Figure 1). Concurrently, we acquired a total of 554 targets associated with the drug ingredients. To standardize the protein target information, we utilized the UniProt database, specifically focusing on the “human” species. After merging the data, 297 duplicate values were removed.
Table 1: Information on the active ingredients of Qianghuo screened in TCMSP. OB, oral bioavailability; DL, drug-likeness.
|
MOL ID |
Name | Compound | OB (%) |
DL |
| MOL001941 | QH1 | Ammidin |
34.55 |
0.22 |
| MOL011962 | QH2 | 6′-Feruloylnodakenin |
32.02 |
0.67 |
| MOL011963 | QH10 | 8-Geranoxy-5-methoxypsoralen |
40.97 |
0.5 |
| MOL011968 | QH11 | coumarin |
33.07 |
0.78 |
| MOL011969 | QH3 | Demethylfuropinnarin |
41.31 |
0.21 |
| MOL011971 | QH12 | Diversoside |
67.57 |
0.31 |
| MOL011975 | QH13 | Notoptol |
62.97 |
0.48 |
| MOL001951 | QH4 | Bergaptin |
41.73 |
0.42 |
| MOL001956 | QH5 | Cnidilin |
32.69 |
0.28 |
| MOL000359 | QH6 | Sitosterol |
36.91 |
0.75 |
| MOL004792 | QH14 | Nodakenin |
57.12 |
0.69 |
| MOL000358 | QH15 | beta-Sitosterol |
36.91 |
0.75 |
| MOL001942 | QH7 | Isoimperatorin |
45.46 |
0.23 |
| MOL002644 | QH8 | Phellopterin |
40.19 |
0.28 |
| MOL002881 | QH9 | Diosmetin |
31.14 |
0.27 |

Figure 1: Compound structural formula.
Leukemia-related Targets
Through the Genecards, TTD, and DisGNET databases, we conducted a keyword search for “leukemia”. We obtained 31 targets from the TTD database, 3507 targets from the Genecards database based on empirical screening with correlations greater than 1.5, and 2111 targets from the DisGeNET database. The targets obtained from the two databases were merged, deduplicated, corrected, and integrated using UniProt. These results constitute the final set of targets related to leukemia for subsequent analysis, totaling 4301 targets (Figure 2).
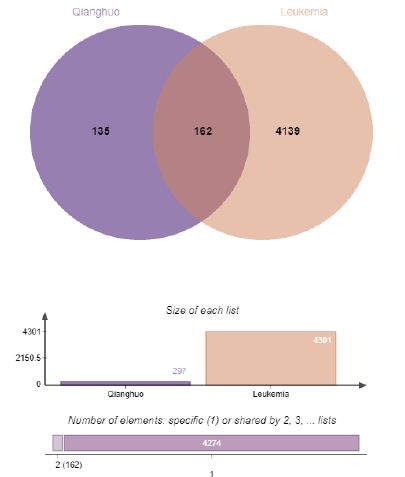
Figure 2: Venn diagram of the intersection targets between Qianghuo and leukemia.
Construction of PPI Network and Screening of Core Proteins
The intersection of the target genes related to the chemical components of Qianghuo and the leukemia target genes was obtained through Venny 2.1.0 and a venn diagram was generated. In the diagram, blue represents the target points of the active ingredients in Qianghuo, totaling 297, while red represents the targets of leukemia, totalling 4301. The overlapping area in the middle of blue and red represents the intersection target, totalling 162, as illustrated in Figure 2. The resulting 162 intersection genes were submitted to the STRING database to analyze the PPI network, as depicted in the Figure 3. The graph comprises 161 nodes and 1982 edges, with an average node degree of 24.6 and a local clustering coefficient of 0.565. Among them, according to the degree value, the top five targets are AKT1, EGFR, SRC, CASP3 and HSP90AA1, in Table 2. Similarly, the top five active ingredients in Qianghuo are Diosmetin, Cnidilin, Bergaptin, and Ammidin and Phellopterin in Table 3. The AKT1, EGFR, SRC, and CASP3 are the hub nodes of the network and can be regarded as potential key targets for the treatment of leukemia by Qianghuo. Akt, a serine/threonine kinase expressed in three isoforms (Akt1, Akt2, and Akt3), acts as the main effector of the PI3K signalling pathway. Its numerous substrates regulate various cellular processes such as protein synthesis, cell cycle progression, cell survival, apoptosis, angiogenesis [16].

Figure 3: Protein interaction map.
Table 2: The top five targets in Drug-ingredient-intersection target-disease network.
|
Gene |
Degree |
|
AKT1 |
100 |
|
EGFR |
89 |
|
SRC |
89 |
|
CASP3 |
84 |
|
HSP90AA1 |
81 |
Table 3: The top five compounds in Drug-ingredient-intersection target-disease network.
|
Compounds name |
Degree |
|
Diosmetin |
58 |
|
Cnidilin |
52 |
|
Bergaptin |
43 |
|
Ammidin |
41 |
|
Phellopterin |
38 |
The PI3K-Akt-mTOR pathway, aberrantly upregulated in cancers including AML, plays a critical role in intracellular signalling [17]. Daniela Bres Sanin et al. [18] demonstrated that targeting the PI3K/Akt/mTOR pathway at different levels can effectively combat cancer cells, leading to the development of dual inhibitors that show efficacy against T-ALL cell lines and primary patient samples. The strong cytotoxic effect of targeting the PI3K/Akt/mTOR pathway demonstrates promising potential for the treatment of T-cell acute lymphoblastic leukemia (T-ALL). Epidermal growth factor receptor (EGFR), a member of the receptor tyrosine kinase (TK) family, is widely expressed in various cancers, including breast cancer, lung cancer, esophageal cancer, and head and neck cancer. EGFR and its family members play pivotal roles in complex signaling cascades that regulate tumor cell growth, differentiation, adhesion, migration, and survival. Consequently, they have emerged as promising candidates for cancer treatment [19]. SRC, a proto-oncogene tyrosine protein kinase, belongs to the non-receptor protein tyrosine kinase family [20]. This family is involved in a plethora of crucial cellular functions essential for maintaining cellular homeostasis, including cell cycle progression, motility, proliferation, differentiation, and survival. Small tyrosine kinase inhibitors have been indicated for the treatment of certain blood malignancies, including CML [21]. Zhang He et al. [22] discovered that SRC tyrosine kinase inhibitors exert an anti-tumor cell role by regulating EGFR and its downstream signaling pathways, which significantly inhibits tumor cell proliferation and infiltration. This finding provides valuable insights for the development of drugs for leukemia treatment. Based on previous research, it has demonstrated potential in leukemia treatment by influencing multiple proven effective targets. These findings suggest that these ingredients play a crucial role in the treatment of leukemia.
Construct Traditional Chinese Medicine-Active Ingredients-Target-Disease and Active Ingredients-Target-Pathway Network Diagram
Import the intersection genes into CytoScape 3.8.0 software to construct a drug-component-target-disease network diagram, as shown in Figure 4. Each goal in the diagram can correspond to one or more components, and the same component can be associated with multiple goals. This shows that Qianghuo has multi-component and multi-target properties in treating leukemia. At the same time, we also constructed a component-target-pathway network diagram, as shown in Figure 5. The top five genes by degree are shown in Table 4. The top five pathways are PI3K-Akt signaling pathway, Human papillomavirus infection, MicroRNAs in cancer and Chemical carcinog, Enesis – reactive oxygen species, Proteoglycans in cancer (Table 5). The above results indicate that the PI3K-Akt signaling pathway is one of the important ways to explore the mechanism of proteoglycans in the treatment of leukemia.

Figure 4: Construction of “herb-active ingredient-target-disease” network.

Figure 5: Construct a network diagram of Qianghuo’s active ingredients-genes-pathway. In the figure, orange ovals represent genes, light green squares represent signaling pathways, and blue squares represent active components.
Table 4: The top five targets in Drug-ingredient-intersection target pathway network.
|
Gene |
Degree |
| PIK3CA |
15 |
| PIK3CB |
15 |
| PIK3CD |
15 |
| MTOR |
12 |
| MAP2K1 |
11 |
Table 5: The top five compounds in Drug-ingredient-intersection target pathway network.
|
Pathway |
Degree |
| PI3K-Akt signaling pathway |
34 |
| Human papillomavirus infection |
29 |
| MicroRNAs in cancer |
26 |
| Chemical carcinogenesis – reactive oxygen species |
25 |
| Proteoglycans in cancer |
24 |
In the figure, the red inverted triangle represents the disease, the blue square represents the intersection target, the green ellipse represents the active ingredient of traditional Chinese medicine, and the pink triangle represents traditional Chinese medicine.
GO and KEGG Enrichment Analysis Results
To elucidate the potential mechanism of action of Qianghuo in the treatment of leukemia, the 162 common targets were imported into the DAVID database for KEGG and GO analysis. The KEGG pathway analysis revealed enrichment in 154 pathways, with prominent pathways including Pathways in cancer, PI3K-Akt signalling pathway (Figure 6), and Human papillomavirus infection (Figure 7A).
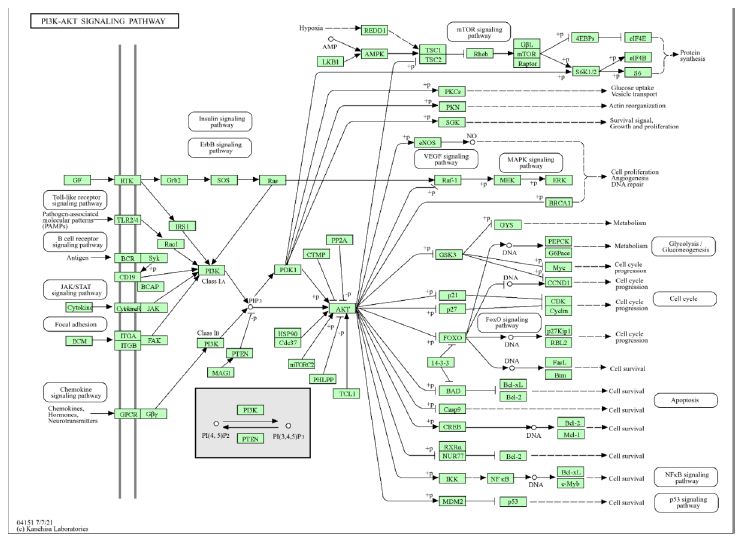
Figure 6: The PI3K-AKT signaling pathway that is the target of Qianghuo treatment of leukemia.
Concurrently, 870 GO biological processes were identified, with 152 related to molecular functions (MF). The top three significantly enriched MF terms were “protein binding,” “ATP binding,” and “protein serine/threonine/tyrosine kinase activity.” Additionally, 634 GO biological processes were linked to biological processes (BP), with the top three enriched terms being “signal transduction,” “protein autophosphorylation,” and “negative regulation of apoptotic process.” Furthermore, 84 GO terms were associated with cellular components (CC), with the top three enriched terms being “cytosol,” “cytoplasm,” and “plasma membrane” (Figure 7B). Among them, the PI3K-AKT pathway has been proven to play an important role in the occurrence and development of leukemia and is a key target for the treatment of leukemia [23]. The PI3K/AKT pathway is involved in cell growth, proliferation, and differentiation, and is often activated in cancer due to various mechanisms such as dysregulation of intrinsic components, mutations in upstream receptors, and/or abnormalities in other interacting signaling molecules [24,25]. Gan and others [26] found that overexpression of miR-217 can inhibit the abnormal activation of the PI3K/Akt pathway by targeting the expression of Akt3, thereby inhibiting cell proliferation, promoting tumor cell apoptosis, and enhancing the sensitivity of doxorubicin to AML. Veronika Ecker et al. [27] reported that the PI3K/AKT pathway can promote the oxidative metabolism pathway in CLL through mar AKTI overexpression, genetic phosphatase SH2-containing-inositol-5′- phosphatase-1 (SHIP1) deletion, or through SHIPI inhibition are activated, leading to the formation of toxic ROS levels. These events ultimately lead to the accumulation of toxic reactive oxygen species (ROS). These findings suggest that the anti-leukemia effect of Qianghuo may be related to multiple signaling pathways, including the PI3K/AKT pathway and oxidative metabolism pathway. Understanding these molecular mechanisms could provide valuable insights into Qianghuo’s efficacy in leukemia treatment.

Figure 7: GO and KEGG enrichment analysis in overlapping targets. (A) KEGG enrichment was consist of the top 10 pathways following the gene ratio. (B) The three sections from left to right were the top five main pathway bubble plots of BP, CC, MF. GO, Gene Ontology; KEGG, Kyoto Encyclopedia of Genes and Genomes; BP, biological process; CC, cellular component; MF, molecular function.
Molecular Docking Results
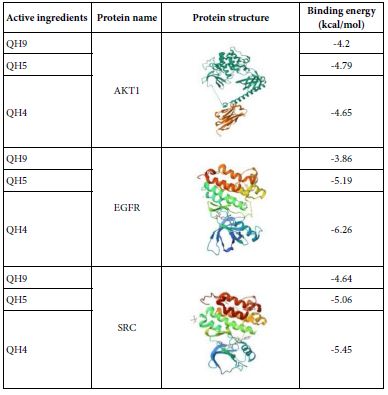
Molecular docking was performed for the hub active ingredients Diosmetin, Cnidilin, Bergaptin, and Ammidin with the core targets AKT1, EGFR, SRC, and CASP3 respectively. Part of the molecular docking results are shown in the Table 7, while the detailed docking results for ligands and receptors are provided in Table 6. In molecular docking, a lower the binding energy indicates a more stable conformation, and a binding energy less than negative five indicates that the ligand and receptor can spontaneously combine. The molecular docking results were consistent with the network pharmacology-related screening results, thereby further validating the reliability of the predicted targets from network pharmacology through molecular docking.
Table 6: Molecular docking visualization of the hub ingredients and hub targets of Qianghuo tablets for the treatment of leukemia.
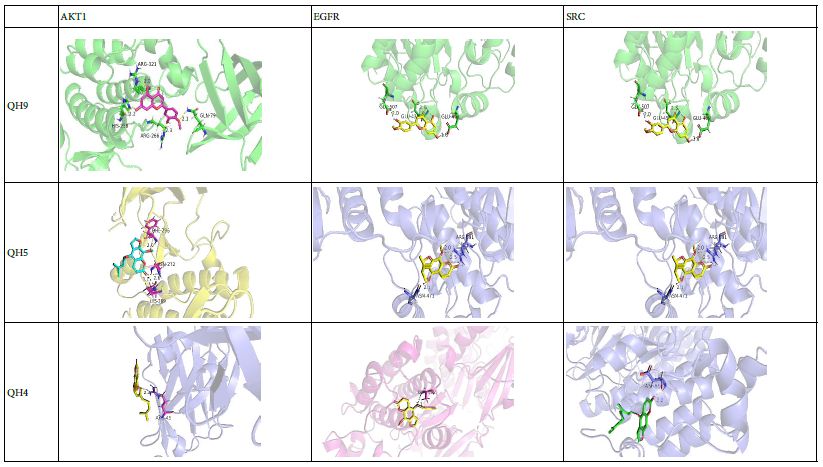
Table 7: Molecular docking results of core targets.
Conclusion
In summary, this study used network pharmacology methods to analyze the mechanism of action of Qianghuo in the treatment of leukemia. It was found that the core active ingredients of Qianghuo in the treatment of leukemia are Cnidilin, Bergaptin, Ammidin, and Diosmetin. By stimulating the cancer signalling pathway, PI3K-Akt signalling pathway, Human cytomegalovirus infection and other pathways, it affects AKT1, EGFR, SRC, and CASP3, and other targets. It plays a role in complex signalling cascades that resist cancer cell proliferation and regulate tumor cell growth, differentiation, adhesion, and survival, thereby effectively inhibiting the infiltration of cancer cells and promoting the recovery of leukemia patients. At the same time, The targeted disease network shows that Qianghuo not only has a good therapeutic effect on cancer, but also has a good effect on other diseases such as rheumatoid, nutritional and metabolic diseases, indicating that different diseases can be treated with the same prescription. Compared with western medicine, which has severe side effects and high cost in treating leukemia, Qianghuo has absolute advantages in treating leukemia with low side effects and low price. In short, we provide an entry point into the mechanism of action of Qianghuo in the treatment of leukemia, and also provide a scientific basis for the treatment of leukemia by Qianghuo. In the next step, in vitro and in vivo experimental verification will be conducted to clarify the specific mechanism of Qianghuo in treating leukemia.
Funding
This work was supported by the Hubei Science and Technology Program (2022CFB781), the Hubei University of Science and Technology (Doctoral Startup Fund Project 170562), and Hubei Provincial Department of Education scientific research guidance project (B2022189).
Conflict of Interest
The authors have no conflicts of interest to declare.
References
- Liu X, Ye F, Wu J, How B, Li W, et al. (2015) Signaling proteins and pathways affected by flavonoids in leukemia cells. Nutr Cancer 67: 238-249. [crossref]
- Nikkila A, Erme S, Arvela H, Holmgren O, Raitanen J, et al. (1975) Background radiation and childhood leukemia: A nationwide register-based case-control study. Int J Cancer 139: 1975-1982. [crossref]
- Tong J, Qin L, Cao Y, Li J, Zhang J, et al. (2012) Environmental radon exposure and childhood leukemia. J Toxicol Environ Health B Crit Rev 2012; 15: 332-347. [crossref]
- Morton LM, Dores GM, Schonfeld SJ, Linet MS, Sigel BS, et al. (2019) Association of Chemotherapy for Solid Tumors With Development of Therapy-Related Myelodysplastic Syndrome or Acute Myeloid Leukemia in the Modern Era. JAMA Oncol 5: 318325. [crossref]
- Tang K, Du S, Wang Q, Zhang Y, Song H (2020) Traditional Chinese medicine targeting cancer stem cells as an alternative treatment for hepatocellular carcinoma. J Integr Med 18: 196-202. [crossref]
- Qi F, Zhao L, Zhou A, Zhang B, Li A, et al. (2015) The advantages of using traditional Chinese medicine as an adjunctive therapy in the whole course of cancer treatment instead of only terminal stage of cancer. Biosci Trends 2015; 9: 16-34. [crossref]
- Wu M, Tang L, Chen Y, Shu L, Xu Y, et al. (2024) Systematic characterization of the chemical constituents in vitro and in vivo of Qianghuo by UPLC-Q-TOF-MS/MS. Fitoterapia.172: 105758. [crossref]
- Zhang R, Zhu X, Bai H, Ning K (2019) Network Pharmacology Databases for Traditional Chinese Medicine: Review and Assessment. Front Pharmacol 10: 123. [crossref]
- Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, et al. (2003) Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13: 2498-2504. [crossref]
- Forli S, Huey R, Pique ME, Sanner MF, Goodsell DS, et al. (2016) Computational protein-ligand docking and virtual drug screening with the AutoDock suite. Nat Protoc 11: 905-919. [crossref]
- Yuan S, Chan HCS, Hu Z (2017) Using PyMOL as a platform for computational drug design. WIREs Computational Molecular Science 7: 2.
- Tao Y, Li W, Yang J, Xue T, Wang Y, et al. (2023) Exploring underlying mechanism of artesunate in treatment of acute myeloid leukemia using network pharmacology and molecular docking. Clin Transl Oncol 25: 2427-2437. [crossref]
- Zhou Z, Chen B, Chen S, Lin M, Chen Y, et al. (2020) Applications of Network Pharmacology in Traditional Chinese Medicine Research. Evid Based Complement Alternat Med.2020: 1646905. [crossref]
- Yang C, Gao H, Yang F, Wang Y (2019) Effects of quinol and isoimperatorin on the inhibition of lipopolysaccharide-induced proliferation of rat fibroblastlike synoviocytes. Pharm Res 38: 621-626+634.
- Chang Y, Chen H, Chen K, Chen K, Chang K, et al. (2016) Investigation of the inhibitors of histone-lysine N-methyltransferase SETD2 for acute lymphoblastic leukaemia from traditional Chinese medicine. SAR QSAR Environ Res 27: 589-608. [crossref]
- Kharas MG, Okabe R, Ganis JJ, Gozo M, Khandan T, et al. (2010) Constitutively active AKT depletes hematopoietic stem cells and induces leukemia in mice. Blood 115: 1406-1415. [crossref]
- Nepstad I, Hatfield KJ, Grønningsæter IS, Reikvam H (2020) The PI3K-Akt-mTOR Signaling Pathway in Human Acute Myeloid Leukemia (AML) Cells. Int J Mol Sci 21: 2907. [crossref]
- Daniela B, Camilla E, Francesca R, Giovanna T, Francesca C, et al. (2012) Harnessing the PI3K/Akt/mTOR pathway in T-cell acute lymphoblastic leukemia: eliminating activity by targeting at different levels. Oncotarget.3: 811-823. [crossref]
- Seshacharyulu P, Ponnusamy MP, Haridas D, Jain M, Ganti AK, et al. (2012) Targeting the EGFR signaling pathway in cancer therapy. Expert Opin Ther Targets.16: 15-31.
- Li S, Nguyen TT, Ung TT, Sah DK, Park SY, et al. (2022) Piperine Attenuates Lithocholic Acid-Stimulated Interleukin-8 by Suppressing Src/EGFR and Reactive Oxygen Species in Human Colorectal Cancer Cells. Antioxidants 11: 530. [crossref]
- Rivera-Torres J, San José E (2019) Src Tyrosine Kinase Inhibitors: New Perspectives on Their Immune, Antiviral, and Senotherapeutic Potential. Front Pharmacol 10: 1011.
- Dorenkamp M, Müller JP, Shanmuganathan KS, Schulten H, Müller N, et al. (2018) Hyperglycaemia-induced methylglyoxal accumulation potentiates VEGF resistance of diabetic monocytes through the aberrant activation of tyrosine phosphatase SHP-2/SRC kinase signalling axis. Sci Rep 8: 14684.
- Alzahrani AS (2019) PI3K/Akt/mTOR inhibitors in cancer: At the bench and bedside. Semin Cancer Biol 59: 125-132. [crossref]
- Liu X, Ye F, Wu Josephine, How Brian, Li W, et al. (2015) Signaling proteins and pathways affected by flavonoids in leukemia cells. Nutr cancer 67: 238-249. [crossref]
- Abrams SL, Ruvolo PP, Ruvolo VR, Ligresti G, Martelli AM, et al. (2017) Targeting signaling and apoptotic pathways involved in chemotherapeutic drug-resistance of hematopoietic cells. Oncotarget 8: 76525-76557. [crossref]
- Gan D, Wu J, Zhou M, Chen W, Jiang W (2024) miR-217 targets the PI3K/Akt pathway to enhance doxorubicin sensitivity in acute myeloid leukemia. Chinese Journal of Experimental Hematology 32: 39-44. [crossref]
- Ecker V, Stumpf M, Brandmeier L, Neumayer T, Pfeuffer L, et al. (2021) Targeted PI3K/AKT-hyperactivation induces cell death in chronic lymphocytic leukemia. Nat Commun 12: 35.