DOI: 10.31038/IDT.2025623
Abstract
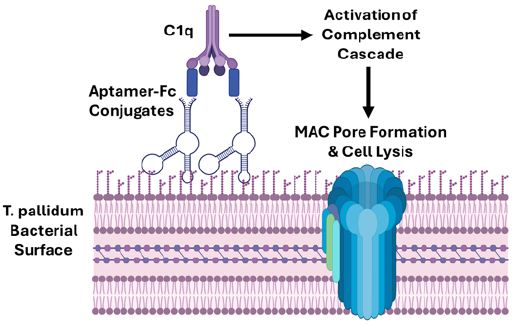
This work extends our recent aptamer-Fc conjugate syphilis opsonin report in this journal (doi.org/10.31038/IDT.2025613) that demonstrated DNA aptamers can be used to couple to the murine immune system and bring about the destruction of Treponema pallidum (causative agent of syphilis) by significantly enhancing macrophage phagocytosis. In the present report, the same anti-Treponema denticola DNA aptamer developed by Park et al. (2015) and designated Td-3 that was used for opsonization was verified to bind a ~ 95 kD surface marker on T. pallidum by aptamer-based Western blotting. The 3’ biotinylated Td-3 aptamer was then bound in a 4: 1 molar ratio to a streptavidin-murine Fc conjugate and in the presence of mouse serum the aptamer-biotin-streptavidin-Fc conjugate killed significant levels of T. pallidum (up to 59%) overnight as assessed by spectrofluorometric analysis using the Invitrogen BacLight™ Live/Dead assay. Thus, this report further expands the potential of aptamer-Fc conjugates as potential artificial “antibody” immunotherapeutics that induce complement-mediated lysis (CML) and may couple to the conjugate’s already documented opsonin effects in the battle against emerging antibiotic-resistant syphilis.
Keywords
Aptamer, Complement, Fc fragment, Lysis, Syphilis, Treponema pallidum
Introduction
The ancient sexually transmitted and congenital disease syphilis persists even today and is increasing in modern society with an estimated 8 million new cases per year in adults between 15 and 49 years of age [1-4]. Despite the continued good efficacy of single dose injectable benzathine penicillin for adults [2,5], benzathine penicillin supply shortages [5], the lack of a vaccine [6], and increasing antibiotic resistance [1], has enabled Treponema pallidum to continue killing about 220,000 people worldwide each year including infants and adults [4]. In particular, the CDC has reported a 79% increase in syphilis cases between 2018 and 2022 with Lieberman et al. [7] reporting that 99.2% of new syphilis cases in North America from 2017 to 2023 were resistant to azithromycin (the alternative antibiotic for patients with penicillin allergy). These numbers sound the alarm for alternative approaches to dealing with this potentially slow, insidious and painful killer pathogen.
The principle of rapid microbial evolution predicts that bacteria such as T. pallidum can and will eventually overcome any and all small molecule antibiotics. Thus, researchers have turned to alternative approaches involving passive immunity through mechanisms such as complement-mediated lysis (CML) and enhanced phagocytosis (opsonization) both of which are known to be at least somewhat effective against T. pallidum [8-15] to eventually treat emerging antibiotic-resistant syphilis. Unfortunately, development and production of humanized monoclonal antibodies for passive CML or opsonization therapy of syphilis or other bacterial diseases would be an expensive proposition. Thus, Bruno et al. previously explored the potentially much less expensive use of DNA aptamer-C1q and -Fc conjugates for CML [16,17] and opsonization [18] of pathogenic bacteria. Indeed, Bruno et al. recently reported in this journal (DOI: 10.31038/IDT.2025613) that an aptamer-biotin-streptavidin-Fc conjugate opsonin performed at least half as well as a commercial polyclonal anti-T. pallidum antibody for enhancement of phagocytosis against this pathogen. In addition, the late Nobel prize laureate for PCR, Dr. Kary Mullis, developed a similar, but less direct, concept with alpha-gal sugar (galactose-α-1,3-galactose) conjugation to antibacterial aptamers (known as “alphamers”) to attract the innate 1% circulating anti-alpha-gal antibodies in human serum [19] which later led to the founding and continued alphamer work of Centauri Therapeutics Ltd. in the UK.
This history and therapeutic needs have brought us to the present report of our efforts to utilize the same aptamer DNA sequence (designated Td-3) developed by Park et al. [20] against a related species T. denticola in an effort to produce aptamer-Fc conjugates capable of CML and opsonization of T. pallidum both in vitro and perhaps eventually in vivo. The following is a summary of our group’s latest work using Park et al.’s Td-3 aptamer from Table 1 to bind a roughly 95 kD surface marker on T. pallidum and when in conjunction with the Fc fragment of murine IgG to couple with the complement system in murine serum to kill up to 59% of T. pallidum overnight in vitro.
Table 1: T. denticola DNA Aptamer Sequences Published by Park et al. [20].

Note: Yellow highlighted regions are identical or nearly identical between the Td-3, -6 and -8 sequences.
Materials and Methods
Treponema pallidum and Bacterial Culture
T. pallidum was purchased from American Type Culture Collection (ATCC; Manassas, VA) as Treponema phagedenis (ex Brumpt) Smibert, Kazan 8 strain (ATCC No. 27087) in lyophilized form. The freeze- dried culture was rehydrated in a 10 ml Oral Treponema Enrichment Broth (OTEB) tube from Anaerobe Systems Inc. (Morgan Hill, CA; Cat. No. AS-603) at 37˚C in a slowly rotating closed incubator with a microaerophilic environment induced by Becton Dickinson Gas Pak™ pouches (Cat. No. 260680). T. pallidum cultures were capped and passaged weekly by 1: 10 dilution of the previous passage in OTEB.
T. denticola Aptamer and Western blotting
Table 1 lists the eight published Park et al. T. denticola aptamer DNA sequences which were purchased from Integrated DNA Technologies (IDT; Coralville, IA) and tested by spectrofluorometry for binding to live T. pallidum cells in our previous recent work (doi.org/10.31038/IDT.2025613) which showed that Td-3 was the consistent top performer that exhibited a high degree of homology with Td-6 and Td-8 (highlighted in Table 1). In the case of Western blotting and CML experiments, we utilized a 3’-biotin-poly A 20mer spacer-Td-3 aptamer DNA oligonucleotide to provide extra distance between the Fc tail of IgG and the T. pallidum bacterial surface. All aptamer sequences were rehydrated in sterile phosphate buffered saline (PBS) without calcium or magnesium and used in fluorescence spectroscopy, microscopy or other experiments as discussed later.
For aptamer-based Western blotting, fresh 3-5 day OTEB cultures of T. pallidum (~ 109 cells) were pelleted at 13,000 X g for 5 minutes and the OTEB culture medium was gently siphoned off. Cells were washed once by resuspension of the bacterial pellet in calcium and magnesium-free PBS (pH 7.2) at room temperature with a second centrifugation in PBS. The pellet was then resuspended in 1 ml of cold 1.5 M MgCl2 and the suspension was stored in a refrigerator overnight. The MgCl2 chaotrope disrupts hydrogen bonding with water and liberates proteins from and in the bacterial surface. Cellular debris was removed by one final pelleting for 5 min at 13,000 X g. Twenty µl of this cell extract was added to 6 µl of 4X LDS NuPAGE™ loading buffer containing beta-mercaptoethanol and samples were heated at 95˚C for 2 minutes. Samples were then loaded into 4-12% gradient Invitrogen Bolt™ Bis-Tris mini polyacrylamide gels and run in cold 1X Bolt™ MES SDS running buffer at 125V for ~ 60 min. Ten µl of Invitrogen PageRuler™ Plus (Cat. No. 26619) prestained molecular weight markers were loaded into the far left lane. After running the gel an identical sister gel was stained with Simply Blue™ rapid Coomassie blue stain followed by rinsing in 100 ml of deionized water and photography. Proteins from the other identical gel were transferred to a PVDF membrane using standard cold transfer buffer containing 20% methanol and transfer at 4 miliAmps overnight.
Chemiluminescent Western blots on the PVDF were developed by first blocking in 10 ml of SuperBlock® from Thermo Fisher for 1 hour at room temperature with gentle mixing. The PVDF membrane was cut into 3 identical segments and exposed to the three different 100 µM biotinylated Td aptamers separately for 1 hour with gentle mixing, followed by 5 rinses in 10 ml of PBS plus 0.1% Tween 20 (PBST) buffer for 5 min per wash. Membranes were then exposed to 10 ml of a 1: 10,000 dilution of Invitrogen streptavidin-alkaline phosphatase conjugate (2 mg/ml stock; Cat. No. 434322) for 1 hour at room temperature, followed by decanting and 5 more rinses in 10 ml of PBST wash buffer and development with Bio-Rad Laboratories Immunostar-AP® substrate (Cat. No. 1705018) and imaging of bands on X-ray film for 10-60 sec.
Streptavidin-Fc Conjugate Formation
In our previous related publication (doi.org/10.31038/ IDT.2025613) we described covalent conjugation of streptavidin (SAv) from Thermo Fisher (Invitrogen Cat. No. 434301) with a murine IgG Fc fragment (Rockland Immunochemicals, Cat. No. 010-0103) using the bifunctional linker BS3 (bis(sulfosuccinimidyl)suberate) from Thermo Scientific (Cat. No. A39266). Successful conjugation was verified by an electrophoretic polyacrylamide gel band shift assay in that recent publication and the same conjugate was used for CML experiments reported herein.
In Vitro CML Experiments and BacLight™ Live/Dead Spectrofluorometric Assay
It is not possible to grow colonies of T. pallidum on the surface of agar spread plates. Therefore, we could not use colony counting to assess T. pallidum viability. Instead we used the popular Invitrogen BacLight™ Live/Dead assay which is based on the cell-permeant green- fluorescing SYTO 9 and red-fluorescing propidium iodide nucleic acid dyes. When alone, SYTO 9 will label all live and dead cells regardless of cell wall and plasma membrane integrity. Propidium iodide (another nucleic acid dye) can only penetrate bacterial cells having damaged cell walls and membranes (i.e., dead cells) where it outcompetes SYTO 9 and leads to red fluorescing cells. The Live/Dead assay has been used broadly and validated on Treponema species previously [21].
For all CML experiments 50 µl of 3-5 day OTEB T. pallidum cultures was added to sterile microfuge tubes with or without 100 µl or 100 µg of the 3’-biotinylated Td-3 aptamer plus 500 µl of frozen and thawed mouse serum (Rockland Immunochemicals Inc., Cat. No. D1080-00-150, Lot No. 43805) and a variable amount of premixed 4: 1 molar ratio of 3’-biotinylated-poly A spacer-Td-3 aptamer plus streptavidin (SAV)-murine Fc BS3 covalent conjugate (~ 1 mg/ml in PBS) as described in our previous opsonin publication. For the variable amounts of the 4: 1 aptamer-biotin-SAv-Fc conjugate 1X (50 µl), 2X (100 µl) and 5X (250 µl) of conjugate were added to different microfuge tubes and calcium and magnesium-free sterile PBS was used to equalize all volumes to 1 ml across each tube. Tubes were mixed briefly, capped and incubated at 37˚C overnight in a microaerophilic environment created by a gas pack in a ziplock bag.
The next morning, tubes were centrifuged at 13,000 X g for 5 min and supernates were aspirated. The cell pellets were resuspended in 1 ml of room temperature PBS without calcium or magnesium. Equal amounts of 3.34 mM SYTO 9 and 20 mM propidium iodide from the Invitrogen Baclight™ Live/Dead kit were mixed together in DMSO. The 1 ml of bacterial cell suspension was added to 2 additional ml of PBS in plastic cuvettes along with 9 µl of the SYTO 9/propidium iodide mixed solution with gentle mixing and cuvettes were incubated at room temperature in the dark for 15 mins. Thereafter, individual cuvette fluorescence spectra were acquired using a Cary Varian Eclipse™ spectrofluorometer with 470 nm excitation 5 nm slits and 700 V photomultiplier tube (PMT) setting for emissions spanning 490 or 500 to 750 nm. The spectral results were validated by fluorescence microscopy using a Thermo Fisher/EVOS FLoid microscope at 400X magnification with two different filter cubes as shown in Figures 2B and 2C.
Results
Figure 1 validates Td-3 and Td-6 aptamer binding to surface molecules extracted from the surface of live T. pallidum cells in the vicinity of ~ 72 to 95 kD with the Td-3 aptamer appearing to bind a ~95 kD moiety (red box and arrow in Figure 1). Td-6 appears to possibly bind a surface molecule of lower molecular weight perhaps closer to 72 kD while the Td-2 aptamer does not appear to bind anything at all. These results correlate well with our recent opsonin- related publication in this journal.
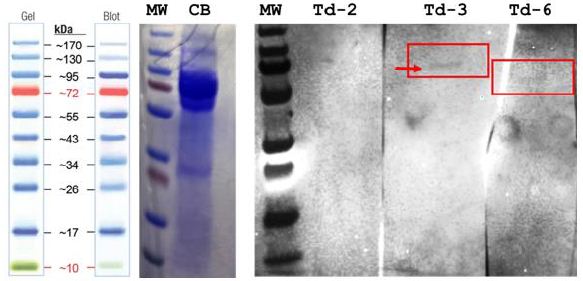
Figure 1: Results of the aptamer Western blots for the Td-2, Td-3 and Td-6 aptamers from Table 1 when used to probe 1.5M MgCl2 T. pallidum live cell extracts demonstrating that Td-3 binds a ~95 kD surface marker, while Td-6 binds a surface molecule closer to 72 kD and Td-2 does not appear to bind. These results correlate with the spectrofluorometric binding results presented in our recent aptamer-Fc opsonin paper in this same journal (doi.org/10.31038/IDT.2025613).
Figure 2 verifies both macroscopically and microscopically that the BacLight™ Live/Dead fluorescence assay stains live T. pallidum 100% green and alcohol-killed T. pallidum 100% red. Unfortunately, due to the limited 400X magnifying power of the available microscope (i.e., 1,000X is needed to do a differential fluorescence count) and clumping of the bacteria seen in Figures 2B and 2C, individual bacterial cell counts were not possible using the available microscope.
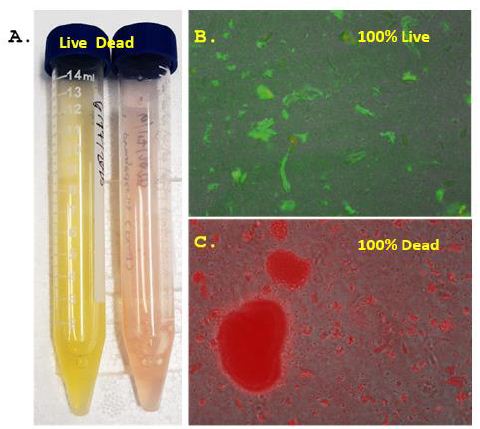
Figure 2: Macro (panel A) and microscopic (400X in panels B and C) results showing the differential green SYTO 9 fluorescent staining of 100% live 3 day T. pallidum culture and red propidium iodide fluorescent staining of 100% dead alcohol-treated T. pallidum cells using the Invitrogen BacLight™ Live/Dead assay kit.
There is not much difference between groups A-C in the red (propidium iodide) region of the spectrum beyond 600 nm in any of the three spectrofluorometry trials (Figures 3-5), suggesting that we would not have seen much of a difference in the numbers of red- fluorescing cells by fluorescence microscopy in any event. However, as show in Figures 3-5, one can estimate cell death percentages by proportion of the fully live cell peaks for the A or B control treatment group spectral traces at 520 nm divided by the peak for the full test aptamer-biotin-SAv-Fc conjugate-treated groups and then subtracting the percent viable cells in the full test C group from 100% to obtain the percent dead cells (i.e., 100% – (100 x group C peak height at 520 nm/the taller of peak A or B height)). Using this formula, we obtained approximately 19.7%, 46% and 59% dead cells for the 1X, 2X and 5X 4: 1 aptamer-biotin to SAv-Fc conjugate concentration treatment groups respectively. The 5X amount (250 µl) of the conjugate is the practical volume limit for these in vitro experiments, thus the limit of killing for the aptamer-Fc conjugate approach is probably about 59% in the full test group C. Test group A was designed to assess the level of killing by mouse serum itself in the alternative pathway [8] while group B was designed to be a negative control to rule out killing of the bacteria by the aptamers themselves. Both groups A and B gave consistently high viabilities in all three conjugate dose trials while the full test group C demonstrated consistent concentration-dependent killing from 19.7% to 59% for the 1X to 5X conjugate doses in vitro.
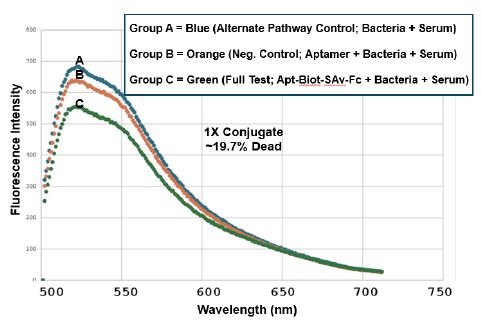
Figure 3: Trial 1 CML spectrofluorometric results using the 1X aptamer-Fc conjugate dose to achieve ~19.7% killing of T. pallidum as assessed by the BacLight™ Lived/Dead assay. Groups A and B were alternate complement pathway and aptamer only negative controls respectively while group C was the full test group in which the greatest level of killing by the aptamer-Fc conjugate plus the other complement components in murine serum was hypothesized to occur by activating the classical complement pathway. The green trace validates that group C has the lowest viability especially in the green region of the spectrum (500 to 550 nm emissions).
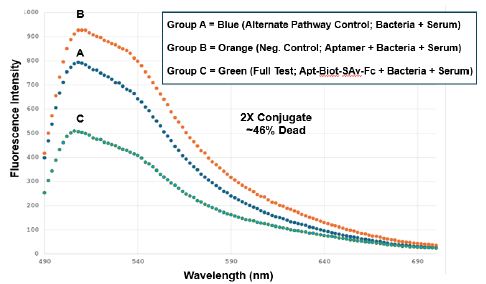
Figure 4: Trial 2 CML spectrofluorometric results using the 2X aptamer-Fc conjugate dose to achieve ~46% killing of T. pallidum as assessed by the BacLight™ Lived/Dead assay. Groups A-C are the same treatment groups as described for Figure 3.
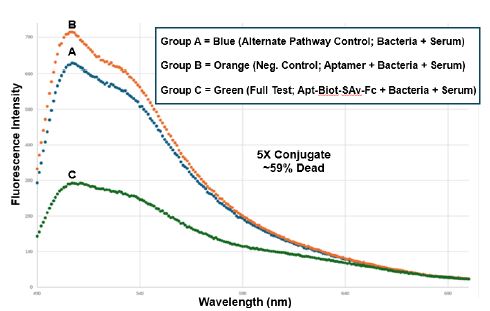
Figure 5: Trial 3 CML spectrofluorometric results using the 5X aptamer-Fc conjugate dose to achieve ~59% killing of T. pallidum as assessed by the BacLight™ Lived/Dead assay. Groups A-C are the same treatment groups as described for Figure 3. The 5X dose represents the practical volume limit for these CML experiments.
Discussion and Conclusions
The present report extends previous publications on aptamer-Fc or -C1q conjugate killing of Gram negative or complement-sensitive bacteria by CML [16,17] to include T. pallidum which is known to be sensitive to antibody-mediated CML [8]. We were hoping to kill nearly 100% of T. pallidum cells via aptamer-Fc-induced CML. However, 59% is a respectable level of killing once the volume limit is hit. And, this is a promising preliminary report especially when coupled to our previous Td-3 aptamer-Fc opsonin publication (doi.org/10.31038/ IDT.2025613) that suggests the remaining 41% of viable T. pallidum which might survive CML in vivo could be opsonized, phagocytized and killed by macrophages or other resident phagocytized in vivo using the same aptamer-Fc conjugate (a proverbial “one-two punch” combination).
In summary, the present work again suggests a new class of aptamer-3’-Fc conjugates to emulate monoclonal antibodies and perhaps lower the cost of future passive immunity CML (and opsonin) therapy. If even higher affinity T. pallidum aptamers can be developed, CML and opsonin-based T. pallidum killing and alternative (non- antibiotic) therapy of syphilis might be even more effective in the future. For use in vivo of course, attachment of the Fc to the 3’ end will be key to extending aptamer conjugate pharmacokinetics and serum half-life by slowing kidney clearance and inhibiting serum exonuclease degradation of the aptamer moiety of the Fc conjugate [22-24].
Acknowledgments
Funding was provided by US NIH (NIAID) SBIR Contract No. 75N93025C00010.
References
- Orbe-Orihuela YC, Sánchez-Alemán M, Hernández-Pliego A, Medina-García CV, Vergara-Ortega DN (2022) Syphilis as Re-Emerging Disease, Antibiotic Resistance, and Vulnerable Population: Global Systematic Review and Meta-Analysis 11: 1546. [crossref]
- Peeling RW, Mabey D, Kamb ML, Chen X-S, Radolf JD, et al. (2017) Syphilis. Nature Reviews Disease Primers 3: 17073. [crossref]
- Stamm LV (2016) Syphilis: Re-emergence of an old Microbial cell (Graz, Austria) 3: 363-370. [crossref]
- Ramchandani MS, Cannon CA, Marra CM (2023) Syphilis: A Modern Infectious disease clinics of North America 37: 195-222. [crossref]
- Seghers F, Taylor MM (2024) Securing the supply of benzathine penicillin: a global perspective on risks and mitigation strategies to prevent future Int Health 16: 279-282. [crossref]
- Avila-Nieto C, Pedreno-Lopez N, Mitja O, Clotet B, Blanco J, et (2023) Syphilis vaccine: challenges, controversies and opportunities. Frontiers in immunology 14: 1126170. [crossref]
- Lieberman NAP, Reid TB, Cannon CA, Nunley BE, Berzkalns A, et al. (2024) Near- Universal Resistance to Macrolides of Treponema pallidum in North N Engl J Med 390: 2127-2128. [crossref]
- Fitzgerald TJ (1987) Activation of the classical and alternative pathways of complement by Treponema pallidum subsp. pallidum and Treponema vincentii. Infect Immun 55: 2066-2073. [crossref]
- Yi DY, Xu QY, He Y, Zheng XQ, Yang TC, et al. (2024) Treponema pallidum protein Tp47 induced prostaglandin E2 to inhibit the phagocytosis in human J Eur Acad Dermatol Venereol 38: 1166-1178. [crossref]
- Chen H, Tong M-L, Liu L-L, Lin L-R, Yang T-C (2022) The whole process of macrophage–Treponema pallidum interactions: Opsonic phagocytosis, nonopsonic phagocytosis and active invasion. International Immunopharmacology 107: 108657. [crossref]
- Hawley K, Cruz A, Benjamin S, Vake C, Cervantes J, et al. (2017) IFNγ Enhances CD64-Potentiated Phagocytosis of Treponema pallidum Opsonized with Human Syphilitic Serum by Human Frontiers in immunology 8: 1227. [crossref]
- Marangoni A, Aldini R, Guardigli M, Sambri V, Giacani L, et (2003) Phagocytosis of Treponema pallidum and reactive oxygen species production by isolated rat Kupffer cells. Medical Microbiology and Immunology 192: 183-188. [crossref]
- Lukehart SA, Shaffer JM, Baker-Zander SA (1992) A subpopulation of Treponema pallidum is resistant to phagocytosis: possible mechanism of The Journal of infectious diseases 166: 1449-1453. [crossref]
- Alder JD, Friess L, Tengowski M, Schell RF (1990) Phagocytosis of opsonized Treponema pallidum subsp. pallidum proceeds slowly. Infect Immun 58: 1167-1173. [crossref]
- COCKAYNE A, PENN CW, BAILEY MJ (1986) Surface Properties of Treponema pallidum in Relation to Phagocytosis by Human Polymorphonuclear Leucocytes in vitro. Microbiology 132: 133-141. [crossref]
- Bruno JG, Carrillo MP, Phillips T (2008) In vitro antibacterial effects of antilipopolysaccharide DNA aptamer-C1qrs complexes. Folia microbiologica 53: 295-302. [crossref]
- Bruno JG (2013) A review of therapeutic aptamer conjugates with emphasis on new Pharmaceuticals 6: 340-357. [crossref]
- Bruno JG, Carrillo MP, Crowell R (2009) Preliminary development of DNA aptamer- Fc conjugate opsonins. Journal of biomedical materials research 90: 1152-1161. [crossref]
- Kristian SA, Hwang JH, Hall B, Leire E, Iacomini J, et (2015) Retargeting pre- existing human antibodies to a bacterial pathogen with an alpha-Gal conjugated aptamer. Journal of molecular medicine 93: 619-631. [crossref]
- Park JP, Shin HJ, Park SG, Oh HK, Choi CH, et (2015) Screening and development of DNA aptamers specific to several oral pathogens. Journal of microbiology and biotechnology 25: 393-398. [crossref]
- Rogers S, Honma K, Mang TS (2018) Confocal fluorescence imaging to evaluate the effect of antimicrobial photodynamic therapy depth on gingivalis and T. denticola biofilms. Photodiagnosis and photodynamic therapy 23: 18-24. [crossref]
- Dougan H, Lyster DM, Vo CV, Stafford A, Weitz JI, et (2000) Extending the lifetime of anticoagulant oligodeoxynucleotide aptamers in blood. Nuclear medicine and biology 27: 289-297. [crossref]
- Bruno JG, Crowell R (2008) Selective glutaraldehyde-mediated coupling of proteins to the 3’-adenine terminus of polymerase chain reaction products. Journal of biomolecular techniques: JBT 19: 177-183. [crossref]
- Keefe AD, Pai S, Ellington A (2010) Aptamers as Nat Rev Drug Discov 9: 537-550.